Ploug Group
Our research focus on structural and functional aspects of proteins involved in disease relevant pathways. The ultimate goal of these studies is to elucidate the function of these proteins in vitro and in vivo and to facilitate the translation of these findings into a clinical setting, particularly within the field of oncology and lipid metabolism.
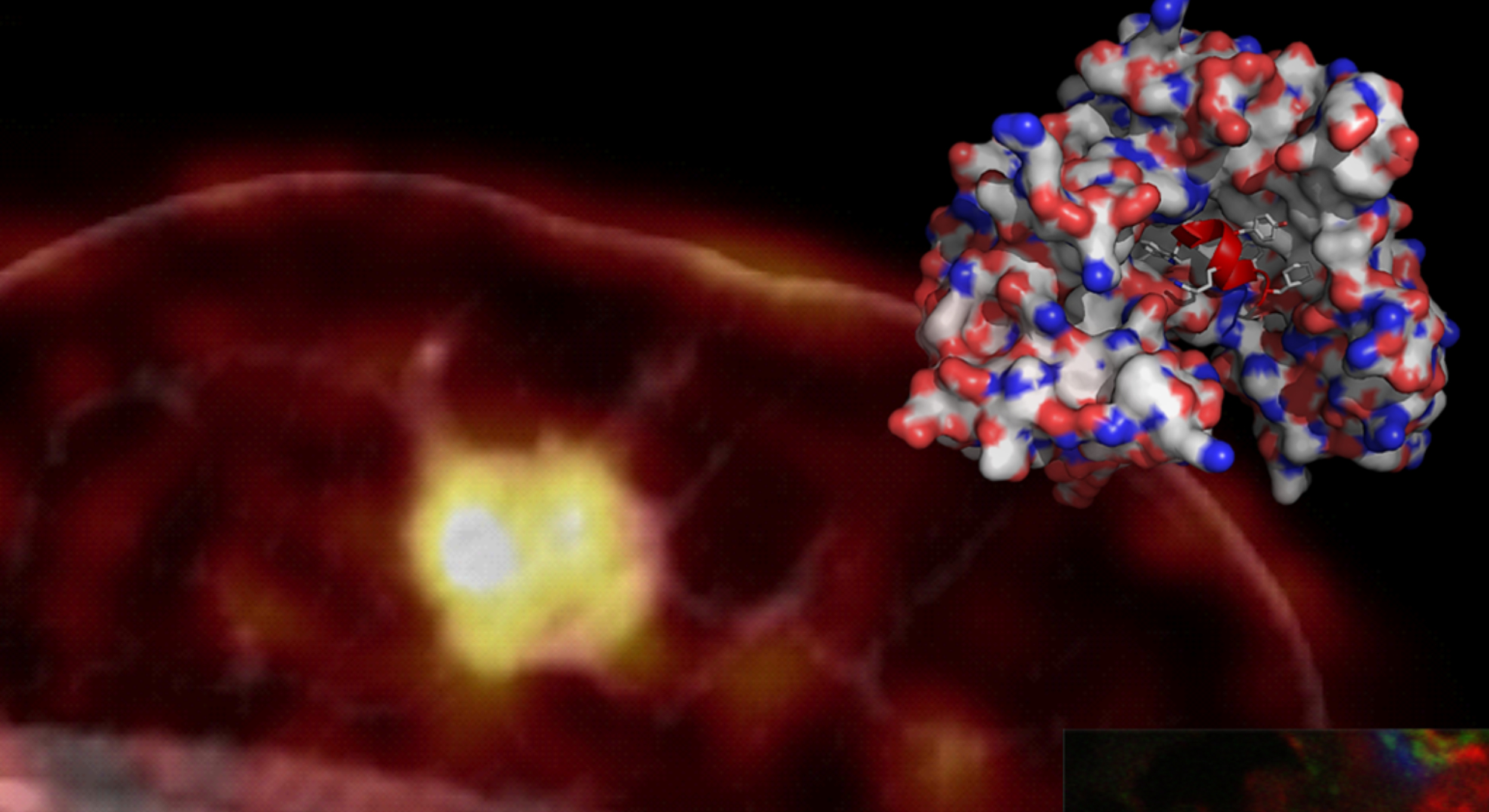
We focus on biochemical and biophysical characterization of disease relevant proteins and their interacting ligands with special emphasis on cancer dissemination and dyslipidemia. We provide detailed understanding on how a given pathway maintains homeostasis in normal physiology and why certain aberrations—such a missense mutations and natural polymorphisms—cause dysregulation in pathophysiology. Ultimately, we seek to use this information in translational pipelines – e.g. our uPAR‐targeting peptide for non‐invasive PET imaging of uPAR‐expression in chronic inflammation and solid cancer.
These discoveries were made by harvesting the full synergy from interactive collaborations.
1: Revised the paradigm for intravascular lipid metabolism—intrinsic protein disorder in GPIHBP1 secures proper compartmentalization of lipoprotein lipase (LPL) and stabilizes the enzyme against spontaneous inactivation.
- Mysling S, et al. (2016) The acidic domain of the endothelial membrane protein GPIHBP1 stabilizes lipoprotein lipase activity by preventing unfolding of its catalytic domain. Elife 5:e12095
- Kristensen KK, et al. (2018) A disordered acidic domain in GPIHBP1 harboring a sulfated tyrosine regulates lipoprotein lipase. Proceedings of the National Academy of Sciences of the United States of America 115(26):E6020
- Song, W, et al. (2022) Electrostatic sheating of lipoprotein lipase is essential for its movement across capillary endothelial cells. J Clin Invest 132 (5):e157500
2: Discovered a new mechanism for LPL regulation—the inhibitor ANGPTL4 catalyzes a cooperative allosteric unfolding leading to the irreversible collapse of LPL’s catalytic triad while the activator APOC2 stabilizes these regions.
- Mysling S, et al. (2016) The angiopoietin-like protein ANGPTL4 catalyzes unfolding of the hydrolase domain in lipoprotein lipase and the endothelial membrane protein GPIHBP1 counteracts this unfolding. Elife 5:20958
- Kristensen KK, et al. (2020) Unfolding of monomeric lipoprotein lipase by ANGPTL4: Insight into the regulation of plasma triglyceride metabolism. Proceedings of the National Academy of Sciences of the United States of America 117(8):4337
- Leth-Espensen KZ, et al. (2021) The instrinsic instability of the hydrolase domain of lipoprotein lipase facilitates its inactivation by ANGPTL4-catalyzed unfolding Proceedings of the National Academy of Sciences of the United States of America 118(12): e2026650118
Kumari A, et al. (2023) Inverse effects of APOC2 and ANGPTL4 on the conformational dynamics of lid-anchoring structures in lipoprotein lipase Proceedings of the National Academy of Sciences of the United States of America 120(18): e2221888120 3: Solved the first crystal structure of LPL.
- Birrane G, et al. (2019) Structure of the lipoprotein lipase-GPIHBP1 complex that mediates plasma triglyceride hydrolysis. Proceedings of the National Academy of Sciences of the United States of America 116(5):1723–1732
4: Discovered a new etiology for acquired chylomicronemia
- Beigneux AP, et al. (2017) Autoantibodies against GPIHBP1 as a Cause of Hypertriglyceridemia. N Engl J Med 376(17):1647–1658
- Lutz J, et al. (2020) Chylomicronemia from GPIHBP1 autoantibodies successfully treated with rituxumab: A case report. Ann Intern Med. doi:7326/L20-0327
5: Developed a peptide probe for non-invasive imaging of solid cancers lesions.
- Persson M, et al. (2015) First-in-human uPAR PET: Imaging of cancer aggressiveness. Theranostics 5(12):1303–1316
- Leth JM & Ploug M (2021) Targeting the urokinase-type plasminogen activator (uPAR) in human diseases with a view to non-invasive imaging and therapeutic intervention. Front Cell Dev Biol 9: PMID: 34490277
Our current research projects have a dual focus: First, we aim at transforming our uPAR-targeted non-invasive PET imaging platform into an optical imaging modality with a view to fluorescence guided intra-operative imaging with improved margin resection. The Holy Grail of successful cancer surgery is to obtain clear resection margins without the need to perform a radical anatomical excision with substantial loss of vital healthy tissue. Second, we will study the molecular mechanisms responsible for ANGPTL-mediated catalysis of LPL inactivation by unfolding. Ultimately, we will used this information to design a stable LPL variant with a view to enzyme replacement therapy in patients with acute dyslipidemia.
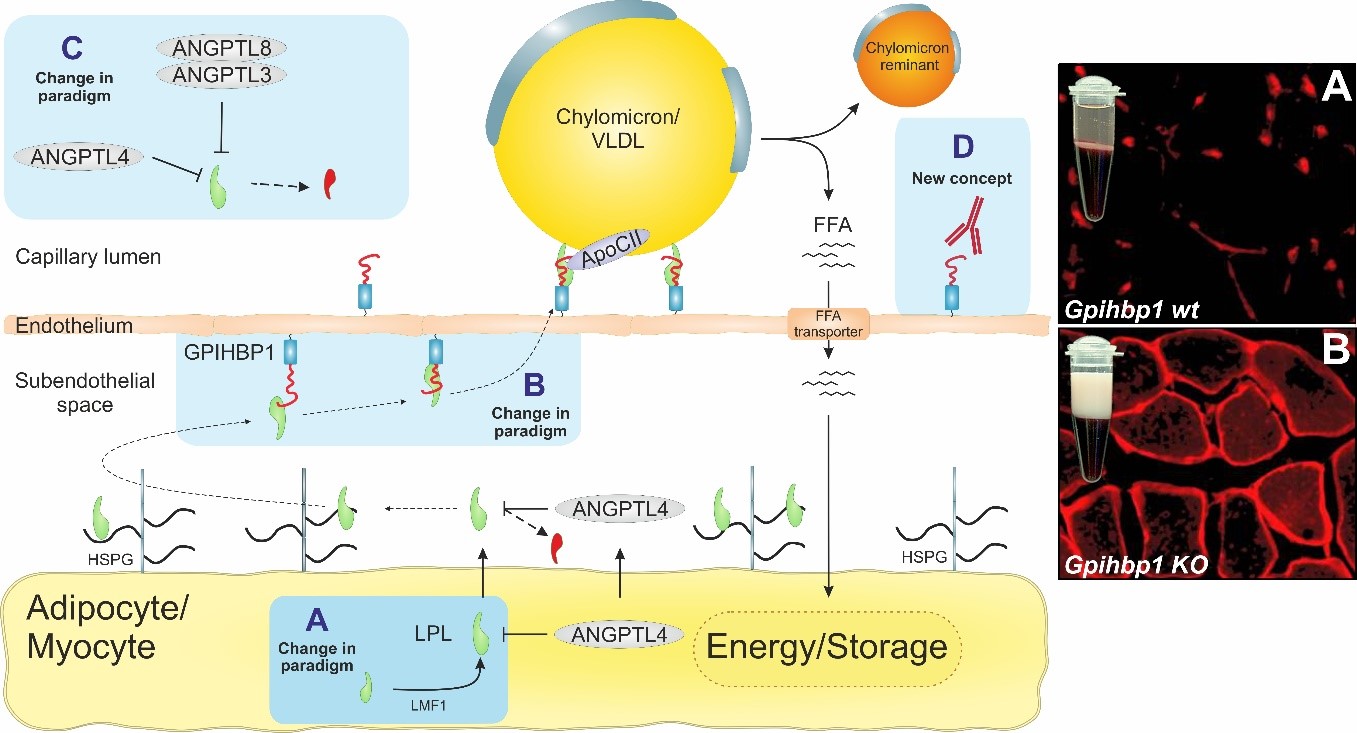
Image 2: Biochemical pathway for intravascular lipid metabolism. Blue squares highlight our research focus areas. Micrographs to the left shows creamy plasma and mislocalization of LPL to the subendothelial spaces when GPIHBP1 is absent or dysfunctional.
To accomplish our research goals, we perform high-end biophysical analyses with surface plasmon resonance (SPR), microscale thermophoresis (MST), hydrogen-deuterium exchange mass spectrometry (HDX-MS), X-ray crystallography, NMR, small angle X-ray scattering (SAXS), and nano-DSF to delineate essential structure-function relationships using highly purified protein preparations produced in house. We have a particularly robust expression system using Drosophila S2 cell, which produces recombinant proteins with uniform glycosylations and are well suited for production of thermolabile proteins. We have a long experience in determining real-time kinetic rate constants with surface plasmon resonance and have used HDX-MS extensively in collaboration with Dr Thomas J. D. Jørgensen.
Selected publications
Kumari A, Grønnemose AL, Kristensen KK, Winther AL, Young SG, Jørgensen TJD, and Ploug M. “Inverse effects of APOC2 and ANGPTL4 on the conformational dynamics of lid-anchoring structures in lipoprotein lipase”
Proc. Natl. Acad. Sci. 120 (2023) PMID: 37094117
Leth-Espensen KZ, Kristensen KK, Kumari A, Winther AL, Young SG, Jørgensen TJD, Ploug M. “The instrinsic instability of the hydrolase domain of lipoprotein lipase facilitates its inactivation by ANGPTL4-catalyzed unfolding”
Proc. Natl. Acad. Sci. 118 (2021) e2026650118
Kristensen, K.K., Leth-Espensen, K.Z., Mertens, H.D.T., Birrane, G., Meiyappan, M., Olivecrona, G., Jørgensen, T.J.D., Young, S.G., and Ploug, M. “Unfolding of monomeric lipoprotein lipase by ANGPTL4: Insight into the regulation of plasma triglyceride metabolism”
Proc. Natl. Acad. Sci. 117 (2020) 4337−4346
Young, S.G., Fong, L.G., Beigneux, A., Allan, C.M., He, C., Nakajima, K., Meiyappan, M., Birrane, G., and Ploug, M. “GPIHBP1 and lipoprotein lipase, partners in plasma triglyceride metabolism”
Cell Metabolism 30 (2019) 51−65
Kristensen, K.K., Midtgaard, S.R., Mysling, S., Korov, O., Hansen, L.B., Skar-Gislinge, N., Kragelund, B.K., Olivecrona, G., Young, S.G., Jørgensen, T.J.D., Fong, L.G., and Ploug, M. “A Disordered Acidic Domain in GPIHBP1 haboring a Sulfated Tyrosine regulates Lipoprotein Lipase”
Proc. Natl. Acad. Sci. 115 (2018) E6020-E6029
Mysling, S., Kristensen, K.K., Larsson, M., Beigneux, A.P., Gårdsvoll, H., Fong, L.G., Bensadoun, A., Jørgensen, T.D., Young, S.G., and Ploug, M. “The acidic domain of the endothelial membrane protein GPIHBP1 stabilizes lipoprotein lipase activity by preventing unfolding of its catalytic domain”
eLife (2016) doi: 10.7554/eLife.12095
Mysling, S., Kristensen, K.K., Larsson, M., Korov, O., Bensadoun, A., Jørgensen, T.J.D., Olivecrona, G., Young, S., and Ploug, M. “The angiopoietin-like protein ANGPTL4 catalyzes unfolding of the hydrolase domain in lipoprotein lipase and the endothelial membrane protein GPIHBP1 counteracts this unfolding”
eLife (2016) doi: 10.77554/eLife.20958

Our studies on intravascular lipolysis are conducted in a close and highly productive collaboration with Dr. Stephen G. Young, Department of Medicine, University of California, Los Angeles, CA (Drs. S. G. Young and A. Beigneux)

The translational studies on non-invasive imaging of uPAR in cancer patients with positron emission tomography (PET) and optical imaging is conducted in close collaboration with Dr. Andreas Kjær Depart Clin Physiol, Nucl Med & PET and Cluster for Molecular Imaging, Rigshospitalet and University of Copenhagen, Denmark (Drs. M. Ploug and A. Kjær)

We have a close collaboration on HDX-MS with Dr. Thomas D.J. Jørgensen BMB, University of Southern Denmark, Odense, Denmark (Drs. S. Mysling, M. Ploug and T.D.J. Jørgensen)