Arnes Group
We are interested in understanding the molecular basis of cellular plasticity in pancreas regeneration and cancer. We collaborate with clinicians and pathologists to bridge our research with patient needs and translate our findings to clinical settings. The long-term goal is to provide better tools to identify early-stage lesions and to develop novel and more specific targets with therapeutic potential.
Pancreatitis increases the risk of pancreatic cancer through not well understood mechanisms. After injury acinar cells repress the expression of genes associated with acinar identity and transiently activate a gene program reminiscent of pancreas development. This mechanism defined as cellular plasticity is a reversible adaptive response enabling cellular proliferation, tissue remodeling, and ultimately recovery from the injury. However, mutations in oncogenes prevent the resolution of cellular plasticity locking the tissue in a proliferative and proinflammatory phase, which generates an oncogenic environment prone to cancer. We envision that pancreatic cancer depends on developmental regulatory elements that become activated during regeneration and can be exploited as novel therapies for this intractable disease.
The main research questions in our lab are: 1) how is cellular plasticity regulated in pancreas regeneration and hijacked in cancer? 2) How oncogenes, transcriptional programs of pancreas development and the tissue microenvironment interplay in cancer initiation and cancer progression? What is the function of pervasively transcribed noncoding RNAs in the transcriptional regulation of cellular plasticity?
Elements of pancreas development become activated upon tissue injury and regulate tumor initiation in genetically engineered mouse models of PDAC
- Developmental transcription factors regulate the specification of heterogenous cellular states in precancerous lesions of cancer
- Cellular plasticity hides cancer cells from the immune system, which is a major impediment in PDAC immunotherapy
- Regulatory element of pancreas development are co-opted by oncogenes and represent Kras-dependencies in cancer initiation
Pancreatic ductal adenocarcinoma is a disease of differentiation and molecular regulators of cellular identity are potent tumor suppressors
- We used an integrative analysis of genomic and clinical data from PDAC tumour samples to define a catalogue of noncoding RNAs associated with genetic traits of pancreatic cancer and associated with clinical outcome.
- Long ncRNAs define molecular subtypes of pancreatic cancer with implications in disease progression and patient stratifications
- molecular subtypes are dynamic, and regulated at least in part by long ncRNAs, opening a window of opportunity to understand and target tumor heterogeneity in pancreatic cancer
- Understand the gene regulation of pancreas regeneration and susceptibility to cancer
- Elucidate the effect of physiological changes associated with aging on cellular plasticity and susceptibility to cancer
- Investigate the molecular mechanisms controlling tumor heterogeneity and resistance to therapy in pancreatic cancer
- Understanding how long noncoding RNAs interact with transcription factors and chromatin modifiers to regulate gene expression and cell fate decision in development and cancer
We use genetic mouse models of PDAC, acinar explants, and human embryonic stem cells to model pancreas development and the early stages of the disease. We combine the results with comparative human data in pathological specimens. We use a combination of gene expression, chromatin accessibility, and single-cell RNA sequencing in time-resolved experiments to define drivers of precancerous lesions and the molecular mechanisms of tumor heterogeneity.
- Mouse models of pancreas regeneration and cancer.
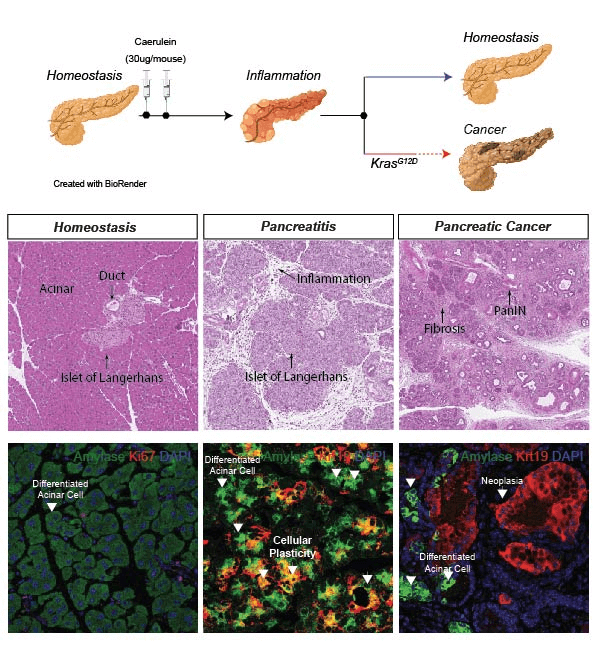
- Modeling pancreatic cancer initiation using human embryonic stem cells
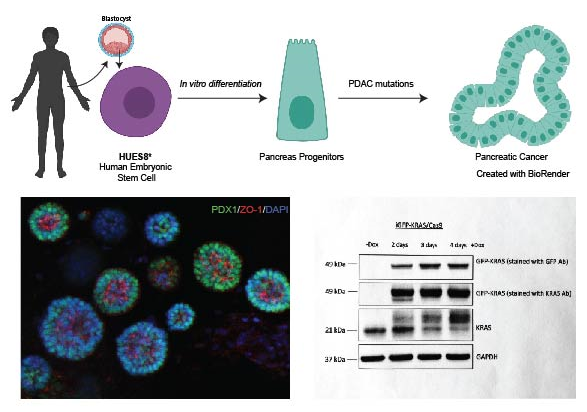
We are using a model of PDAC initiation using an inducible K-RAS activating mutation (K-RASG12V) in human embryonic stem cells. Top figure shows a schematic representation of the model depicting the steps to generate PDAC lesions from human embryonic stem cells. Botton figure shown pancreas progenitors derived from HUES8 grown in 3D. Molecular evidence showing the tight regulation of KrasG12V expression with doxycycline. *Generously provided by Prof. Danwei Huangfu. MSK. New York
- Patient derived organoids
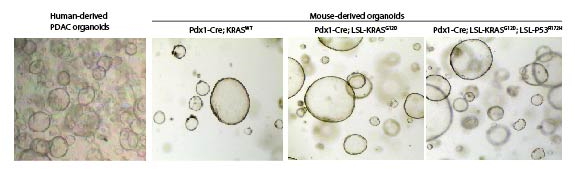
The laboratory is using a combination of human- and mouse-derived organoids to investigate the molecular mechanisms of cellular plasticity in homeostasis and cancer.
- Integrative computational analysis.
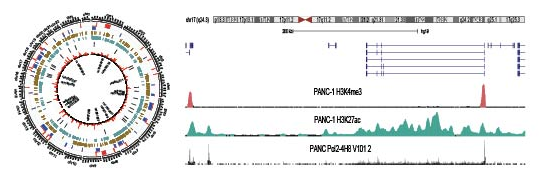
- Rolver, MG., et al. (2025). Tumor microenvironment acidosis favors pancreatic cancer stem cell properties and in vivo metastasis. iScience (accepted).
- Perez-Penco, M., et al. The antitumor activity of TGFβ-specific T cells is dependent on IL-6 signaling. Cell Mol Immunology. 10.1038/s41423-024-01238-7.
- Baldan, J., et al. (2024). Resolution of acinar dedifferentiation regulates tissue remodeling in pancreatic injury and cancer initiation. Gastroenterology. 10.1053/j.gastro.2024.04.031. #
- Høj, K., et al. (2024). Age-related decline in pancreas regeneration is associated with an increased proinflammatory response to injury. Gastro Hep Advances. 10.1016/j.gastha.2024.07.002. #
- Lewinska, M., et al. (2024). Fibroblast-Derived Lysyl Oxidase Increases Oxidative Phosphorylation and Stemness in Cholangiocarcinoma. Gastroenterology. 10.1053/j.gastro.2023.11.302.
- Rafaeva, M., et al. (2023). Fibroblast-derived matrix models desmoplastic properties and forms a prognostic signature in cancer progression. Front Immunol 14, 1154528. 10.3389/fimmu.2023.1154528.
- Thijssen, M. S., Bintz, J. & Arnes, L. In Vitro Silencing of lncRNA Expression Using siRNAs. Methods in molecular biology 2348, 141-156, doi:10.1007/978-1-0716-1581-2_9 (2021).
- Nielsen, S. R. et al. Suppression of tumor-associated neutrophils by lorlatinib attenuates pancreatic cancer growth and improves treatment with immune checkpoint blockade. Nature communications 12, 3414, doi:10.1038/s41467-021-23731-7 (2021).
- Montes, M. & Arnes, L. lncRNAs: potential therapeutic targets and biomarkers for pancreatic cancer? Expert opinion on therapeutic targets 25, 521-528, doi:10.1080/14728222.2021.1938541 (2021).
- Lopez de Maturana, E. et al. A multilayered post-GWAS assessment on genetic susceptibility to pancreatic cancer. Genome Med 13, 15, doi:10.1186/s13073-020-00816-4 (2021).
- Rabadan, R. et al. Identification of relevant genetic alterations in cancer using topological data analysis. Nature communications 11, 3808, doi:10.1038/s41467-020-17659-7 (2020).
- Singer, R. A. et al. The Long Noncoding RNA Paupar Modulates PAX6 Regulatory Activities to Promote Alpha Cell Development and Function. Cell metabolism 30, 1091-1106 e1098, doi:10.1016/j.cmet.2019.09.013 (2019).
- Arnes, L. et al. Comprehensive characterisation of compartment-specific long non-coding RNAs associated with pancreatic ductal adenocarcinoma. Gut 68, 499-511, doi:10.1136/gutjnl-2017-314353 (2019).
- Font-Cunill, B., Arnes, L., Ferrer, J., Sussel, L. & Beucher, A. Long Non-coding RNAs as Local Regulators of Pancreatic Islet Transcription Factor Genes. Frontiers in genetics 9, 524, doi:10.3389/fgene.2018.00524 (2018).
- Arnes, L., Akerman, I., Balderes, D. A., Ferrer, J. & Sussel, L. betalinc1 encodes a long noncoding RNA that regulates islet beta-cell formation and function. Genes & development 30, 502-507, doi:10.1101/gad.273821.115 (2016).
- Arnes, L. & Sussel, L. Epigenetic modifications and long noncoding RNAs influence pancreas development and function. Trends in genetics : TIG 31, 290-299, doi:10.1016/j.tig.2015.02.008 (2015).





