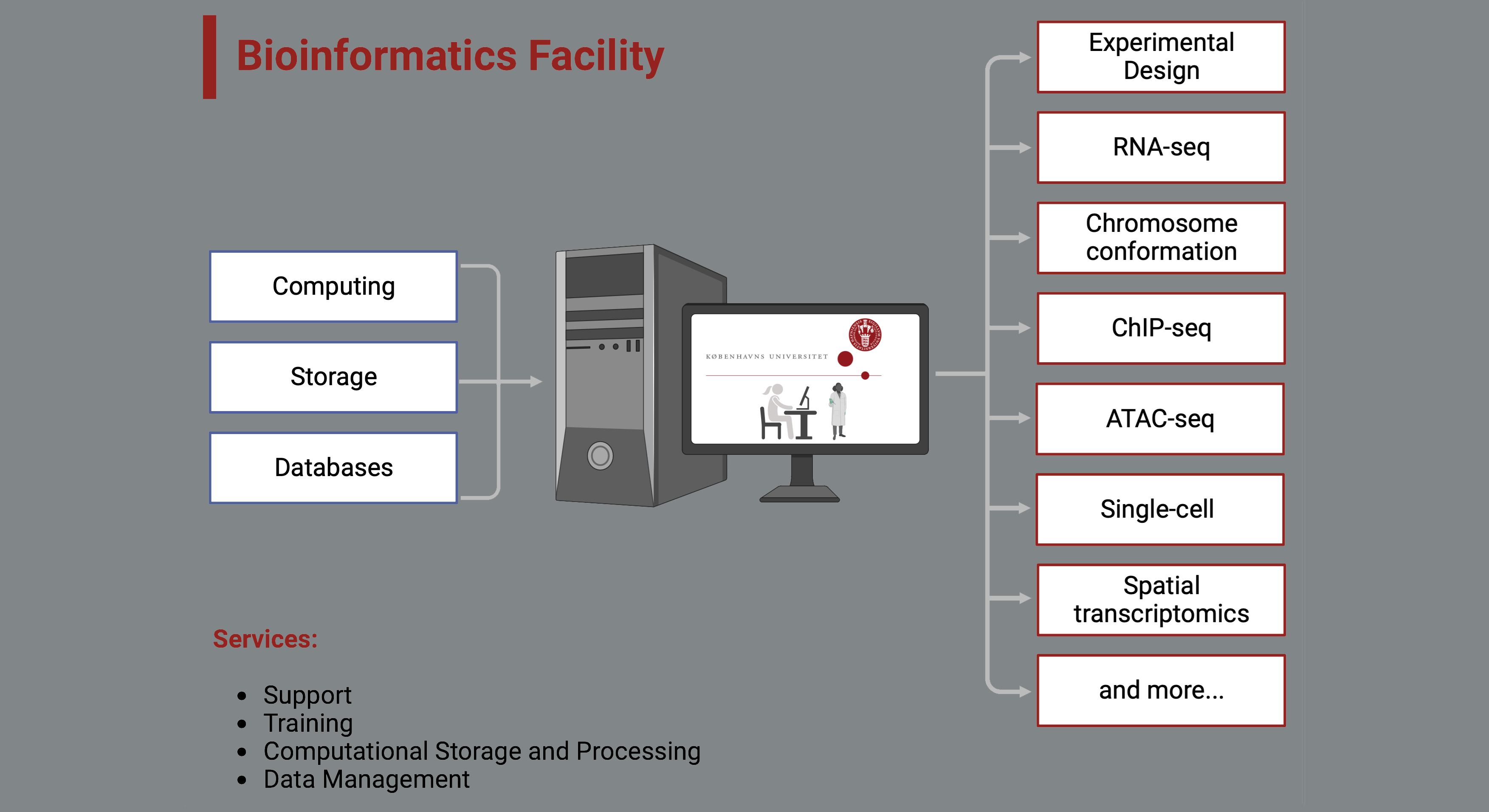
Our services
Our support mainly involves consultation, experimental desing, data analysis, data management, guidance, short & long-term projects and training. Although you are welcome to contact us for other bioinformatic needs.
Multiple services are offered:
1. Bioinformatic analysis services (with a priority to BRIC users)
2. Access to a maintained high-performance computational server (only for BRIC users)
3. Training in bioinformatics (only for BRIC users)
4. And more.
Only for BRIC users:
Access to a computing and storage infrastructure with:
- Automated backup
- Software tools for formatting, analyzing, visualizing, interpreting and comparing biological data.
- Databases and knowledge bases storing biological knowledge.
- Training programs on demand on handling and analyzing data for users at BRIC with diverse backgrounds and experience.
The services below are available to all users, both BRIC users and external users:
You are always welcome to talk to us! We can work together to understand your needs.
No fee is applied to understand your needs
Including a bioinformatician in an early stage of your project can save you time, work, and money.
We adapt to your needs. We can discuss and find together the best way to analyze your data. Also, if you want to analyze the data yourself we can provide support for your bioinformatic analysis with your own code or with existing pipelines.
A standard bioinformatic project would be a project with typically around 6 samples, 3 samples of each condition, where the user aims to perform a differential expression analysis between the two conditions and a functional enrichment analysis. The data types included in such category are: 1. RNA-seq
2. ChIP-seq
3. ATAC-seq
4. Single Cell data
Estimating hours is always challenging; We estimate a workload of 20 to 30 hrs for a standard analysis of simple datasets.
An Example of a Standard RNA-seq dataset analysis:
- Initial meeting / communications
- Reading about the project
- Transfer raw sequencing data
- Quality assessment of the data
- Communication with the user
- Alignment of sequencing reads onto the genome of interest and gene quantification
- Differential gene expression analysis
- Functional enrichment
- Report and Data visualization
- Meetings to discuss results and possible follow-ups
A plus in using our services: We work with optimized pipelines ensuring reproducibility and providing reports.
We aim to be able to offer an End-to-End service for your project. We work in close collaboration with other BRIC core facilities to offer and End-To-End service
Library Preparation - Sequencing - Bioinformatics
Please, Talk to us for more information on how to get the best of us.
-
You can also request our services for Data Management. We can prepare and submit your data to any public database.
-
Don’t hesitate to contact us to help you preparing your manuscript or grant application.