Spatial RNA and protein profiling
At the Bartholin Institute, we are dedicated to advancing the field of spatial omics through cutting-edge technology and innovative research. Our team comprises experts in molecular biology, bioinformatics, and imaging techniques, providing comprehensive services tailored to meet the needs of researchers and institutions.
We offer state-of-the-art facility and resources for GeoMX spatial transcriptomics and proteomics, enabling detailed analysis of tissue samples at unprecedented resolution. Our goal is to support scientific discovery by providing high-quality data and expert guidance throughout your research journey.
We are actively seeking collaborations with researchers, academic institutions, and industry partners. If you are interested in exploring the potential of spatial omics or have a project in mind, we would love to hear from you. Together, we can push the boundaries of knowledge and unlock new insights into biological processes.
GeoMX Digital Spatial Profiler (GeoMX DSP)
The GeoMX Digital Spatial Profiler (DSP) is a cutting-edge platform that enables high-resolution spatial profiling of proteins and RNA within tissue samples. Using ultraviolet (UV) light and barcoded oligonucleotide probes attached to antibodies or RNA targets, GeoMX precisely maps the expression of hundreds of molecules across regions of interest (ROIs) in variety of complex tissues, such as tumors or brain sections.
After selecting specific ROIs, based on morphology markers fluorescent staining, UV light releases the barcoded probes from the tissue, which are then collected and quantified using nCounter or next-generation sequencing (NGS). The technology allows researchers to study molecular interactions in situ while preserving the spatial context, providing valuable insights into tissue heterogeneity and disease pathology.
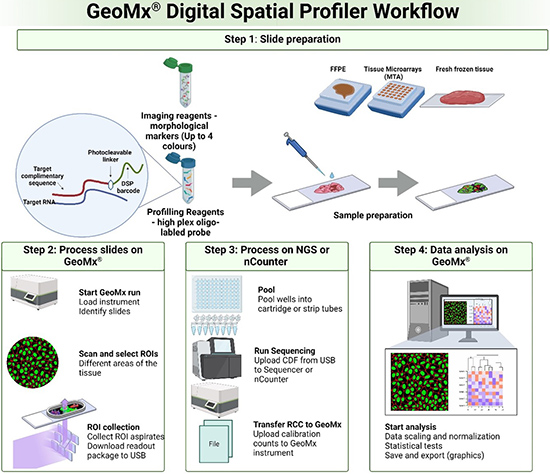
Sample Preparation
Tissue sections are prepared and mounted on slides, then stained with fluorescently labeled antibodies or RNA probes targeting specific proteins or transcripts.
Region of Interest (ROI) Selection
Using the GeoMX platform's high-resolution imaging, specific regions of interest (ROIs) within the tissue are selected, based on morphological markers.
UV Light Cleavage
UV light is directed to the selected ROIs, releasing barcoded oligonucleotide tags attached to the probes, while preserving the tissue architecture.
Barcode Collection & Quantification
The released barcodes are collected and quantified using next-generation sequencing (NGS) or digital counting, providing spatially resolved data for protein or RNA expression.
Data Analysis
The quantified molecular data are analyzed, correlating spatial expression patterns with tissue architecture, disease states, or other biological features.
Consultation
Schedule a consultation with the core facility team to discuss project feasibility, experimental design, and assay selection. Academic and collaborative projects with common publications are prioritized.
Project Proposal
Submit a detailed project proposal outlining objectives, tissue types, RNA or protein profiling, and specific regions of interest (ROIs) to be analyzed.
Sample Submission
Ensure samples are properly prepared and meet GeoMX requirements (e.g., tissue type, thickness, fixation) before submission.
Data Ownership
Data generated from collaborative projects will be shared with the project PI. Any publication or presentation must acknowledge the GeoMX core facility.
Cost Sharing
Collaborators are responsible for costs associated with time used (staff scientist and bls), reagents, sequencing, and data analysis, as agreed upon before project initiation.
Timeline
Projects are processed on a first-come, first-served basis. Estimated timelines will be provided during consultation.
Authorship
Core facility staff contributing substantially to experimental design, data acquisition, or analysis should be considered for co-authorship in publications.
Data Storage
Raw data will be stored for a fixed period (e.g., 6 months) after the project is completed; long-term storage is the responsibility of the collaborator.
Harwood, D.S.L., Pedersen, V., Bager, N.S. et al. Glioblastoma cells increase expression of notch signaling and synaptic genes within infiltrated brain tissue. Nat Commun 15, 7857 (2024).
Tekin H, Lindhardt C, Antvorskov JC, Bager NS, Michaelsen SR, Areškevičiūtė A, Vind JP, Kristensen BW, Josefsen K. Using GeoMx DSP Spatial Proteomics to Investigate Immune Infiltration of NOD Mouse Islet and Exocrine Compartments. Mol Imaging Biol. 2024 Nov 18. doi: 10.1007/s11307-024-01961-7. Epub ahead of print. PMID: 39557779.
Puig-Blasco L, Piotrowski KB, Michaelsen SR, et al. Loss of cancer cell-derived ADAM15 alters the tumor microenvironment in colorectal tumors. Int J Cancer. 2023; 153(12): 2068-2081. doi:10.1002/ijc.34695.
Contact
Mirolyuba Ilieva
mirolyuba.simeonova.ilieva@regionh.dk
Bartholin Institute
Ole Maaløes Vej 5,
buil. 2, 3rd floor, 2.3.19
