Cvejic Group
The mission of my lab is to understand how blood cells develop and how is their development and their ultimate function determined/influenced by their environment and environmental cues. We study how stem and progenitor cells make cell fate choices in the context of niche they are surrounded by. The ultimate aim of the lab is to use this knowledge to develop new ways to treat disease and improve human health.
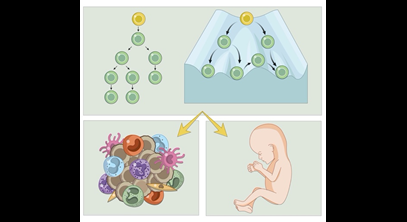
Blood stem cells need to both perpetuate (self-renew) themselves and differentiate into all mature blood cells to maintain blood formation throughout life. Clarifying how haemopoietic stem and progenitor cells (HSPCs) differentiate into diverse cell types is important for understanding how this process is subverted in the generation of blood pathologies. The aim of our group is to decipher how differentiation pathways of HSPCs are influenced by different microenvironments. To achieve that we use state-of-the-art single-cell data generation combined with computational analysis to establish principles of blood lineage differentiation.
With the principal expertise and research interest in the molecular regulation of blood stem cell fate choices our research sits at the intersection of molecular biology, genetics and systems biology and it closely couples experimental approaches and “big” biological data analysis. Some of the main finding from the lab include:
- Changes in motif accessibility in HSCs precedes activation of lineage specific transcriptional programs and their cell fate choice
- The primitive wave of human haematopoiesis likely originates from the hypoblast
- Tens of somatic mutations are accumulated in the first eight weeks of human development and the mutation rate is not constant during early development
We are focusing on the dissection of the heterogeneity of cellular states in the blood system. Our research involves the use of primary human tissues. Currently, we are working with human foetal haematopoietic cells to reveal the dynamics and cellular programmes active during human blood development as well as cancer patient samples to investigating the influence of tumour microenvironment in the context of pathological differentiation of myeloid progenitors.
The results from our studies will advance our understanding of how normal fate decisions are instigated and provide clues for the design of novel therapies for blood pathologies.
Selected publications
Spencer Chapman M, Ranzoni AM, Myers B, Williams N, Coorens THH, Mitchell E, Butler T, Dawson KJ, Hooks Y, Moore L, Nangalia J, Robinson PS, Yoshida K, Hook E, Campbell PJ, Cvejic A (2021). Lineage tracing of human development through somatic mutations. Nature. 2021 Jul;595(7865):85-90.
Ranzoni AM, Tangherloni A, Berest I, Riva SG, Myers B, Strzelecka PM, Xu J, Panada E, Mohorianu I, Zaugg JB, Cvejic A (2020). Integrative Single-Cell RNA-Seq and ATAC-Seq Analysis of Human Developmental Hematopoiesis. Cell Stem Cell. 2020 Dec 15:S1934-5909(20)30553-1.
Hernández PP, Strzelecka PM, Athanasiadis EI, Hall D, Robalo AF, Collins CM, Boudinot P, Levraud JP, Cvejic A (2018). Single-cell transcriptional analysis reveals ILC-like cells in zebrafish. Sci Immunol, 2018 Nov 16;3(29). pii: eaau5265.
Athanasiadis EI, Botthof JG, Andres H, Ferreira L, Lio P, Cvejic A (2017). Single-cell RNA-sequencing uncovers transcriptional states and fate decisions in haematopoiesis. Nat Commun, 2017 Dec 11;8(1):2045.
Macaulay IC, Svensson V, Labalette C, Ferreira L, Hamey F, Voet T, Teichmann S, Cvejic A(2016). Single cell RNA-sequencing reveals a continuous spectrum of differentiation in haematopoietic cells. Cell Reports, 14(4):966-7.



