Service
We are working with the Chromium 10x system and offering:
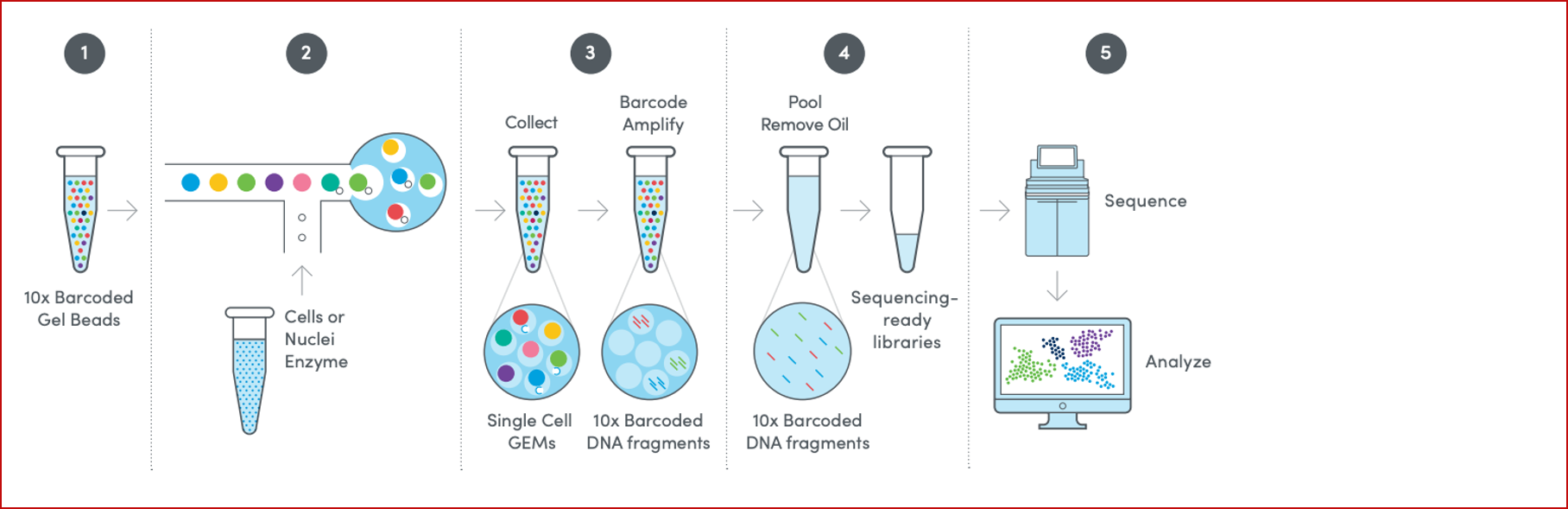
- 10X ‘Next GEM Technology’ for single cell 3′ GEX assay
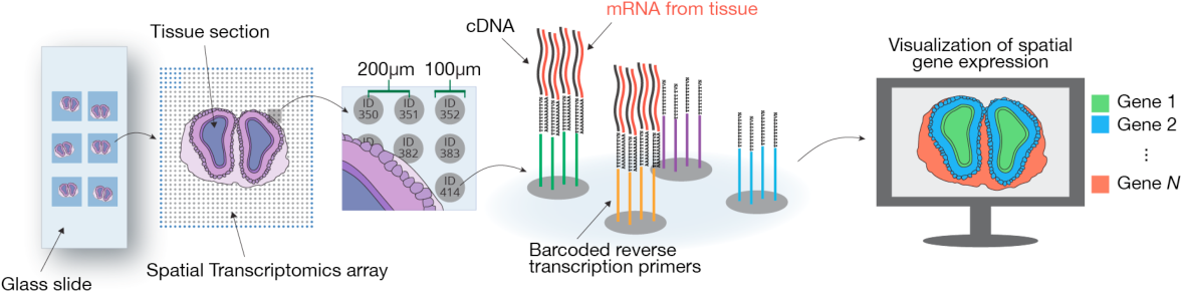
- Visium Spatial Gene Expression
|
At! Our Single cell facility does not provide FACS, sequencing and data analysis. - If you need FACS, please perform it at your Institute or book time at BRIC FACS facility. - For sequencing contact Sequencing Facility at KU or BRIC or other sequencing services around Denmark/the World. - For data analysis contact BRIC Bioinformatics core facility or other Bioinformatics facilities around Denmark/the World. |
- 10x “University”- a great place to start. Here you will find a wealth of training videos, seminars etc. We highly recommend that you watch the videos, relevant for your experiment, before planning any 10x experiments and/or going in the lab
- Workflow 10x Training– presentation in Power Point
- Cell Preparation Guide
- Demonstrated protocols
- Customer Developed Protocols
- 10x Questions and answers
Pay attention to the proper preparation of cells or nuclei from tissue. This is a crucial step of the method. We are not able to do it for you, since different types of cells need different methods of isolation. We take isolated cells from you and work with that solution of cells that you bring. Below you will find general requirements for a sample.
Cell sizes:
There is no minimum cell size. Cells can have a diameter of up to 30 µm. Cells larger than 30 µm increase the risk of clogging Chip and should not be run. As an alternative, it is recommended isolating nuclei from these samples.
10X Chromium Single Cell suspension sample requirements:
- Cell suspensions should always be kept on ice and where possible proceed with cell loading immediately after sample preparation.
- Minimum concentration of 100 cells/ul (700 to 1,200 cells/ul optimal range) in a volume of at least 40ul.
- If at all possible, bring 70ul of single cell suspension (two attempts at chip loading in case of clog plus additional for cell QC). 10 ul from the sample is for the cell counter.
- Single cell suspension should be at least 70% live (Trypan Blue stained) and free of visible debris and doublets. Bothdebris and clamped cellswill have influence on your results and even can clog up a chipduring cell recovering, which makes impossible to finish recovering of cells.
- Recommended cell suspension buffer is PBS/0.04% BSA. Cell suspension buffers should be free of EDTA and Mg++ as well as free of DNAse to be compatible with single-cell assay.
- Cell suspension buffers that can be used:
- 1X PBS + 0.04% BSA
- HBSS + 0.04% BSA
- EMEM + 10% FBS
- DMEM + 10% FBS
- RPMI + 10% FBS
- Hams F12 + 10% FBS
- IMDM + 10% FBS
- DMEM:F12 + 10% FBS
- FACS and 10X:FACS samples are compatible with the 10x Single Cell workflow. FACS is a good method to enrich for specific cell types based on cell surface markers, and to get a clean cell suspension as it also removes the dead cells and debris. If you are using a rare cell type, it is also possible to sort directly into the media that you will use with the 10x Single Cell Master Mix (PBS+0.04% BSA for example). Note: Cell counts from FACS are often overestimated. We strongly recommend recounting cells after flow sorting f.e. under microscope
- How many cells (scRNASeq)
- Approximately 65% loaded cells will be recovered after 10x machine sorting.
- The number of cells per sample is highly dependent on the number of cell types present in the sample and the fractional abundances of the rarest cell types. If you have an idea about the heterogeneity of the sample, you can use the calculator developed by the SATIJA LAB to estimate a minimum number of cells/sample. Link to calculator
- Basically, 700 to 16000 cells per reaction should be loaded to chip to get recommended range 500 to 10,000 recovered cells (check page 24 in the Chromium 10x Protocol that we send you together with this letter).
Briefly: One sample should contain at least 700-16.000 cells (ideally 16000) with concentration 700-1200 cells /ul . Always count cells under microscope (even after FACS) and check for debris and not dissociated cells in the solution.
Book time and make orders of reagents here
Please schedule any 10X single cell experiment at least a week in advance.
We highly advise a consultation prior to experiment scheduling if you are not experienced.