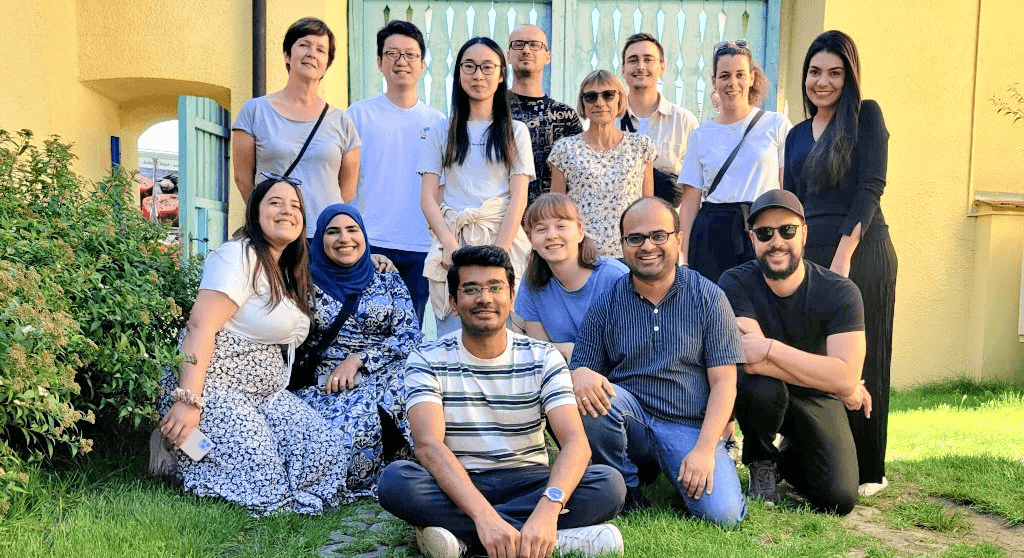
Khodosevich Group
We aim to address the key questions for our understanding of brain function “how such complex organ as the brain is build up during development?” and “what goes wrong in neurodevelopmental disorders?”.
We study mechanisms that are responsible for neuronal specification, positioning and circuit formation during brain development, and how these mechanisms are impaired in neurodevelopmental disorders, such as schizophrenia, epilepsy, autism and others. In particular, we investigate how genetics and environment shape brain development and determine the vulnerability of neurons to developmental disorders.
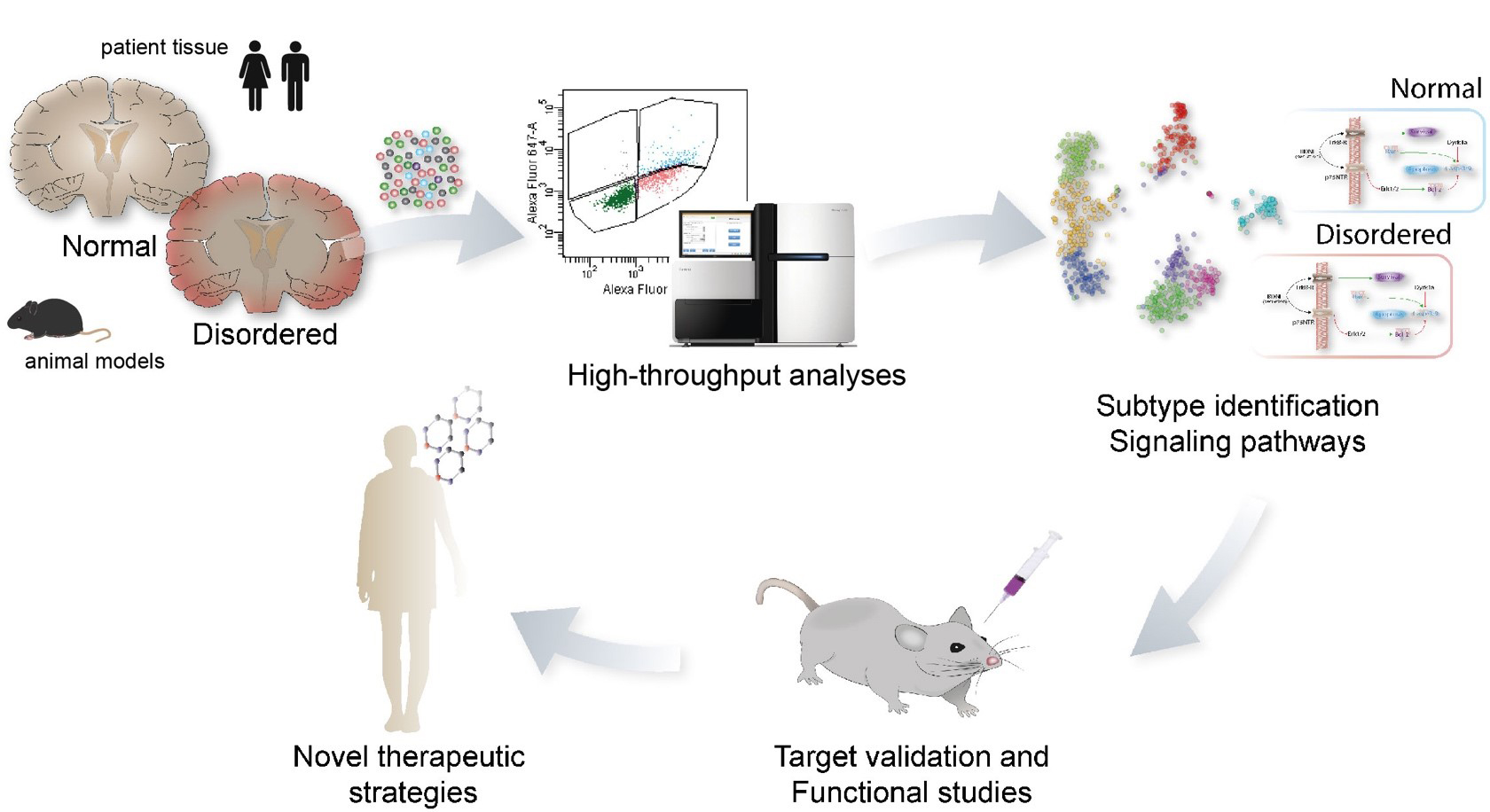
- Response to genetic and environmental stimuli determines the vulnerability of neurons to neurodevelopmental disorders.
- The vulnerability of neurons is highly dependent on developmental period; thus, there are critical periods of development, when different types of neurons are particularly vulnerable, or particularly resilient, and these periods differ for different neuronal types
Read papers: Maternal inflammation has a profound effect on cortical interneuron development in a stage and subtype-specific manner & Identification of Vulnerable Interneuron Subtypes in 15q13.3 Microdeletion Syndrome Using Single-Cell Transcriptomics
- Neuronal types, circuits and gene expression at single-cell resolution that contribute to epilepsy and schizophrenia in human brain.
Read papers: Identification of epilepsy-associated neuronal subtypes and gene expression underlying epileptogenesis & Selective vulnerability of supragranular layer neurons in schizophrenia
- Identification of a pivotal metabolic gene to pair energy production and energy demand in fast-spiking interneurons (the key neurons in neurodevelopmental disorders)
- Specific metabolism of fast-spiking interneurons programs their neuronal maturation, and when impaired leads to neuropsychiatric phenotype
Normal brain development
- Neuronal specification in normal brain.
- Maturation of cortical interneurons.
Neurodevelopmental disorders
- Epilepsy:
- Mechanisms of epileptogenesis in human tissue and animal models.
- Functional characterization of human neurons in epileptic tissue
- Schizophrenia:
- Effects of environmental factors, such as maternal viral infections, inflammation, and pre-term birth on fetal brain development and postnatal brain maturation.
- Effects of schizophrenia-associated genetic mutations on fetal brain development and postnatal brain maturation.
- Formation of neuronal circuits underlying schizophrenia.
- Metabolic impairment in schizophrenia.
Single Cell Omics and Spatial Analysis
We are working to understand brain development and neurodevelopmental disorders at a molecular scale, and how various molecular perturbations alter the brain function. Our research extends from studying disease model rodent brains to human patient brain tissues. We apply single-cell and single-nucleus omics as well as spatial omics methods to profile molecular and spatial alterations in the tissue. Our approach provides a comprehensive insight into the alterations underlying impaired brain function in neurodevelopmental disorders.
Lab
- High-throughput profiling: We use the 10X Genomics Chromium platform. This droplet-based method enables profiling of several thousand of cells or nuclei in a parallel manner and thus permits the exploration of the cellular composition of a given tissue.
- High-resolution profiling: We use Smart-seq2 for high-resolution profiling of single-cells or nuclei. This method provides greater sequencing depth and thus potentially allows detecting subtle molecular alterations in the normal and disordered brain.
- Spatial profiling: We implement spatial analysis based on 10X Genomics Chromium platform allowing us to identify changes in cell composition and spatial location of molecular changes in disordered brains.
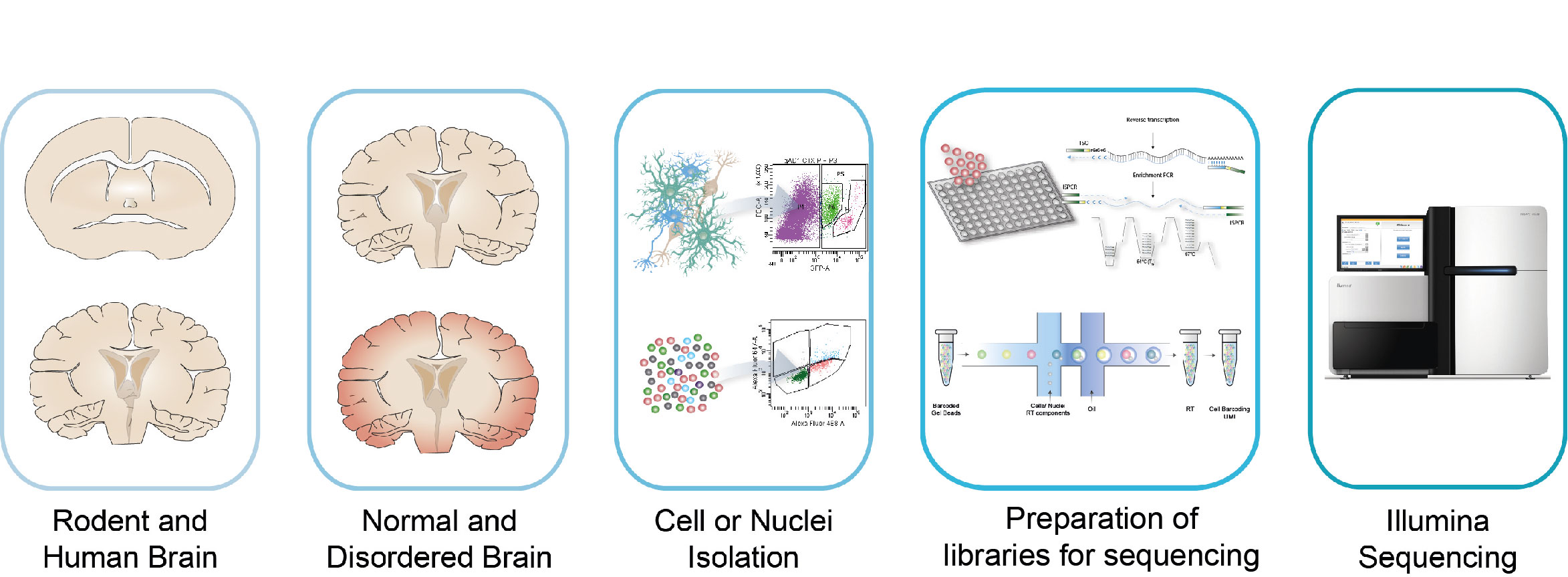
Bioinformatics
Following single-cell RNA sequencing, downstream analysis involves
Identification of differentially expressed genes and pathways
Clustering of the cells according to their gene expression
Visualisation and interactive exploration of the data
Characterisation of the disease-related changes
Integration of several modalities of single cell data (e.g. transcriptome and epigenome), integration with genetics, with spatial data etc.
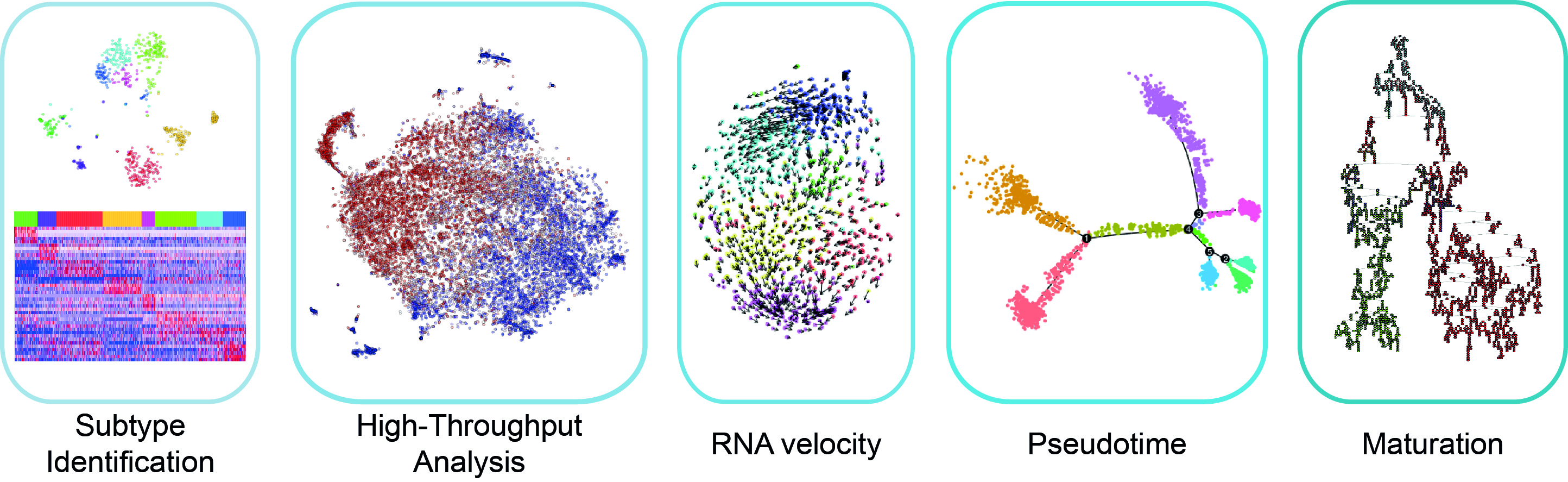
Tools we commonly use for these tasks include the packages Cacoa, Conos, Pagoda 2, velocyto, scanpy and paga, as well as pipelines as dropest, CellRanger, stereoscope and CellPhoneDB.
Selected publications
Experimental papers
Batiuk M#, Tyler T#, Dragicevic K, Mei S, Rydbirk R, Petukhov V, Deviatiiarov R, Sedmak D, Frank E, Feher V, Habek N, Hu Q, Igolkina A, Roszik L, Pfisterer U, Garcia-Gonzalez D, Petanjek Z, Adorjan I, Kharchenko P, Khodosevich K (2022) Upper cortical layer–driven network impairment in schizophrenia. Sci Adv 8, eabn8367
Malwade S, Gasthaus J, Bellardita C, Andelic M, Moric B, Korshunova I, Kiehn O, Vasistha NA, Khodosevich K. (2022) Identification of vulnerable interneuron subtypes in 15q13.3 microdeletion syndrome using single-cell transcriptomics. Biol Psychiatry, 91(8): 727-739
Pfisterer #, Demharter S.#, Petukhov V.#, Meichsner J.#, Thompson J., Batiuk M., Asenjo Martinez A., Vasistha N., Thakur A., Mikkelsen J., Adorjan I., Pinborg L., Pers T., von Engelhardt J., Kharchenko P., Khodosevich K. (2020) Identification of epilepsy-associated neuronal subtypes and gene expression underlying epileptogenesis. Nat Commun, 11(1):1-19
Vasistha N.#, Pardo-Navarro M.#, Gasthaus J., Weijers D., Müller M., García-González D., Malwade S., Korshunova I., Pfisterer U., von Engelhardt J., Hougaard K., Khodosevich K. (2019) Maternal inflammation has a profound effect on cortical interneuron development in a stage and subtype-specific manner. Mol Psychiatry, 10.1038/s41380-019-0539-5
Sanz Morello B., Pfisterer U., Hansen N., Demharter S., Thakur A., Fujii K., Levitskiy S., Montalant A., Korshunova I., Mammen P., Kamenski P., Noguchi S., Aldana Garcia B., Hougaard K., Perrier J., Khodosevich K. (2020) Unique isoform in oxidative phosphorylation machinery supports the function of fast-spiking neurons. EMBO J, 39(18):e105759
Computational papers
Petukhov V#, Igolkina A#, Rydbirk R, Mei S, Christoffersen L, Khodosevich K#, Kharchenko P# (2022). Case-control analysis of single-cell RNA-seq studies. bioRxiv 2022.03.15.484475.
Petukhov V, Xu R, Soldatov RA, Cadinu P, Khodosevich K, Moffitt J, Kharchenko P. (2021) Cell segmentation in imaging-based spatial transcriptomics. Nat Biotech, 40(3): 345-354
Barkas N.#, Petukhov V.#, Nikolaeva D., Lozinsky Y., Demharter S., Khodosevich K., Kharchenko PV. (2019). Joint analysis of heterogeneous single-cell RNA-seq dataset collections. Nat Methods, 2019 Jul 15. doi: 10.1038/s41592-019-0466-z.
Reviews & Perspectives
Khodosevich K, Dragicevic K, Howes O. (2023) Drug targeting in psychiatric disorders – how to overcome the loss in translation? Nat Rev Drug Discovery, s41573-023-00847-7
Khodosevich K, Sellgren CM. (2023) Neurodevelopmental disorders—high-resolution rethinking of disease modeling. Mol Psychiatry 28(1):34-43
In the media
Forskere på KU afslører de specifikke neuron-fejl bag skizofreni
19.10.2022 by Ingeniøren (story also brought in Dagens Medicin and on bric.ku.dk)
Et bestemt gen kobles til autisme, ADHD, skizofreni og andre diagnoser
02.11.2021 by Videnskab.dk
New video: How to build a healthy brain.
Study identifies brain cells most affected by epilepsy and new targets for their treatment
07.10.2020 by Faculty of Health and Medical Sciences
BRIC researchers discover role of metabolism gene in brain dysfunction
04.08.2020 by BRIC
How are psychiatric disorders linked to infections during pregnancy?
30.10.2019 by Faculty of Health and Medical Sciences
View Khodosevich Group on Twitter
Main collaborators
Peter Kharchenko, Harvard Medical School, USA
Lars Pinborg, Rigshospitalet, Denmark
Jean-Francois Perrier, University of Copenhagen, Denmark
Marco Capogna, University of Aarhus, Denmark
Istvan Adorjan, Semmelweis University, Hungary
Tune Pers, University of Copenhagen, Denmark
Jakob von Engelhardt, University of Mainz, Germany
Kazunobu Sawamoto, Nagoya City University, Japan
Kedar Natarajan, University of Southern Denmark, Denmark
Pascal Fries, Ernst Strüngmann Institute, Germany
Susana Aznar, Bispebjerg Hospital, Denmark
Ana Pombo, Max Delbrück Center, Germany
Karin Sørig Hougaard, National Research Center for Work Environment, Denmark








