Theilgaard-Mönch Group
Our research group conducts a patient-centric translational preclinical program aiming at the identification and validation of novel therapeutic strategies eradicating LEUKEMIC STEM CELLS in patients with ACUTE MYELOID LEUKEMIA. In addition, we apply our newly identified human hematopoietic stem cell (HSC) marker to refine the human HSC immunophenotype and explore its clinical potential for production of highly purified stem cell grafts applicable for autologous stem cell transplantation of AML and other cancer patients, as well as gene therapy of patients with monogenic diseases
The Theilgaard-Mönch Group is located at the Finsen Laboratory, a cancer research department at the Finsen Centre at Rigshospitalet, Copenhagen University Hospital.
Acute Myeloid Leukemia (AML) is an aggressive cancer entity with a dismal clinical outcome due to relapse and primary resistance toward standard chemotherapy regimens.
AML is organized as a cellular hierarchy initiated by a rare population of leukemic stem cells (LSCs), which originates from normal hematopoietic stem and progenitor cells (HSPCs) that have acquired molecular and chromosomal driver aberrations propagating cell properties leading to leukemic transformation. Hence, similar to normal HSCs, LSCs have a unique ability to self-renew, and to differentiate and produce a bulk population of more mature AML blast cells. Significantly, LSCs exhibit a slower turnover rate and consequently a high resistance towards chemotherapy as compared to the bulk population of AML cells.
Our research group aims to develop novel therapies combining targeted inhibition of cancer-specific vulnerabilities in AML LSCs with standard therapies to enhance the therapeutic efficacy by eradicating AML at its root. Additionally, we aim to apply our novel HSC marker, which is not expressed by AML cells, to produce purified “tumor-free” stem cell grafts, in order to reduce the risk of relapse and improve clinical outcome of ACUTE MYELOID LEUKEMIA (AML) patients, after high-dose therapy combined with autologous stem cell transplantation.
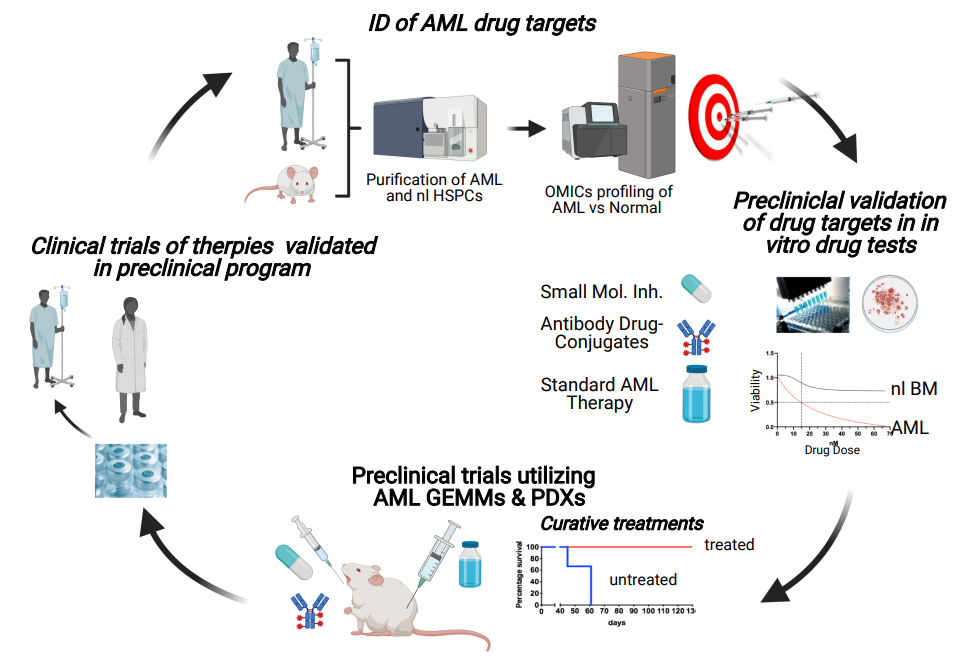
- Demonstration that AML patients whose prognostic driver aberrations emerge in hematopoietic stem cells (HSCs) have significant worse prognosis as compared to patients whose driver aberrations emerge in hematopoietic progenitor cells (HPCs) (Mora-Jensen et al., 2015).
- Functional characterization of the ERG oncogene as a regulator of HEMATOPOIETIC STEM CELL (HSC) maintenance using a conditional Erg knock-out mouse model (Knudsen et al., 2015).
- Identification of aberrant oncogenic signaling and DNA damage response pathway activities in patients with AML (Rapin et al., 2014) and demonstration that combinations of small molecule inhibitors targeting these pathways effectively inhibit survival of primary AML cells in vitro and in vivo (Estruch et al., 2020).
- Identification of a NOVEL HUMAN HSC MARKER (Reckzeh et al., 2018). This marker is expressed on normal human HSCs but not on AML cells. Our novel marker represents the only currently known only currently known surface marker that is conserved on mouse and human HSCs!
- Identification of a novel surface marker that is expressed by LSCs of a genetically defined AML patient subclass as well as cancer cells from patients with other rare hematologic malignancies and solid cancers, but not by normal human hematopoietic stem and progenitor cells (HSPCs) (unpublished data). So far, we have generated a first generation ANTIBOBDY DRUG-CONJUGATE and demonstrated that our ADC targeting this surface molecule effectively eradicates AML in vitro, and in AML xenograft/PDX trials (unpublished data).
NOVEL HSC MARKER – Biology and clinical application:
- We have generated KO mice to investigate the function of our novel HSC marker, which currently is the only known surface marker that is conserved on mouse and human HSCs, as well as during development on fetal liver HSCs, cord blood HSCs, and adult bone marrow HSCs! We are currently exploring the potential of our novel HSC marker to refine the human HSC immunophenotyped allowing for improved purification of human HSCs.
- Given that our novel HSC marker is NOT expressed by AML and other cancer patient cells we are developing a clinical protocol for the production of purified “tumor-free” stem cell grafts for autologous stem cell transplantation, which ultimately holds great potential to reduce the risk of relapse and improve clinical outcome of AML and cancer patients treated with high-dose chemotherapy and autologous stem cell transplantation. Our novel HSC marker also holds great potential for purification of HSCs for gene therapy of monogenic disorders such as severe combined immunodeficiency, transfusion-dependent thalassemia, and sickle cell disease.
AML - Development of novel therapies targeting LSCs:
- Development of novel AML combination therapies: Application of OMICS analyses and in vitro drug tests of primary AML patient cells (i.e. LSCs) for identification and therapeutic targeting of cancer vulnerabilities such as oncogenic signaling, apoptosis, and DNA damage response pathways, in order to ultimately enhance sensitivity and overcome resistance toward chemotherapy. Effective drug combinations from our in vitro tests are validated in preclinical trials using genetically engineered AML mouse model and AML patient-derived xenografts, to identify novel effective multi-drug therapies targeting LSCs for future investigator-initiated trials.
- Development of novel ANTIBOBDY DRUG-CONJUGATE (ADCs) for AML LSC treatment Identification of novel actionable surface antigens for ADC treatment of AML LSCs. So far, we have identified one relevant surface marker and shown that our own first-generation ADCs targeting this novel surface molecule effectively eradicates AML in vitro and in xenograft/PDX trials (unpublished data) without substantial toxicity toward normal HSPCs. As part of a recent biotech program, we are currently developing our own proprietary high-affinity MoAbs to generate a novel improved ADC for treatment of AML LSCs and other cancer entities expression the identified actionable surface marker.
- Multi-color flowcytomery analysis and sorting of HSPC populations
- “Omics analyses - NGS, RNAseq, collaboration on MS – Phosphoproteomics (Jesper V Olsen Lab at CPR, UCPH)
- AML GEMMs (Genetically Engineered Mouse AML Models)
- AML Patient-Derived Xenografts (PDX)
- In vitro drug tests – AML GEMMs and AML patients
- Preclinical drug trials – AML GEMMs and AML PDXs
- Estruch M, Reckzeh K, Vittori C, Centio A, Ali M, Engelhard S, Zhao L, Won KJ, Liu P, Porse BT, Theilgaard-Mönch K. Targeted inhibition of cooperative mutation- and therapy-induced AKT activation in AML effectively enhances response to chemotherapy. Leukemia, 2020 Dec 9. https://doi.org/10.1038/s41375-020-01094-0
- Roy D, Thongjuea S, Theilgaard-Mönch K, Nerlov C. Identification of two distinct pathways of human myelopoiesis Science Immunology (2019):4 (35). https://doi.org/10.1126/sciimmunol.aau7148
- Jakobsen JS, Laursen LG, Schuster MB, Pundhir S, Schoof E, Ge Y, d'Altri T, Vitting-Seerup K, Rapin N, Gentil C, Jendholm J, Theilgaard-Mönch K, Reckzeh K, Bullinger L, Döhner K, Hokland P, Fitzgibbon J, Porse BT. Mutant CEBPA directly drives the expression of the targetable tumor-promoting factor CD73 in AML. Sci Adv. 2019 Jul 10;5(7):eaaw4304. https://advances.sciencemag.org/content/5/7/eaaw4304
- Reckzeh K, Kizilkaya H, Helbo AS, Estruch M, Deslauriers AG, Grover A, Rapin N, Asmar F, Grønbæk K, Porse B, Borregaard N, Vestweber D, Nerlov C, Theilgaard- Mönch K. Human adult HSCs can be discriminated from lineage-committed HPCs by the expression of endomucin. Blood Adv(2018) 2 (13): 1628–1632. DOI: https://doi.org/1182/bloodadvances.2018015743
- Theilgaard-Mönch K. Gut microbiota sustains hematopoiesis. Blood. 2017 Feb 9;129(6):662-663. https://ashpublications.org/blood/article/129/6/662/36313/Gut-microbiota-sustains-hematopoiesis
- Mora-Jensen H, Jendholm J, Rapin N, Andersen MK, Roug AS, Bagger FO, Bullinger L, Winther O, Borregaard N, Porse BT, Theilgaard-Mönch K. Cellular origin of prognostic chromosomal aberrations in AML patients. Leukemia. 2015 Aug;29(8):1785-9. https://doi.org/10.1038/leu.2015.30
- Knudsen KJ, Rehn M, Hasemann MS, Rapin N, Bagger FO, Ohlsson E, Willer A, Frank AK, Søndergaard E, Jendholm J, Thorén L, Lee J, Rak J, Theilgaard-Mönch K and Porse BT (shared last authorship). ERG promotes the maintenance of hematopoietic stem cells by restricting their differentiation. Genes Dev. 2015 Sep 15;29(18):1915-29. http://genesdev.cshlp.org/content/29/18/1915
- Rapin N, Bagger FO, Jendholm J, Mora-Jensen H, Krogh A, Kohlmann A, Thiede C, Borregaard N, Bullinger L, Winther O, Theilgaard-Mönch K and Porse BT (shared last authorship). Comparing cancer vs normal gene expression profiles identifies new disease entities and common transcriptional programs in AML patients. Blood. 2014 Feb 6;123(6):894-904. https://ashpublications.org/blood/article/123/6/894/32581/Comparing-cancer-vs-normal-gene-expression
- Theilgaard-Mönch K, Boultwood J, Ferrari S, Giannopoulos K, Hernandez-Rivas JM, Kohlmann A, Morgan M, Porse B, Tagliafico E, Zwaan CM, Wainscoat J, Van den Heuvel-Eibrink MM, Mills K, Bullinger L. Gene expression profiling in MDS and AML: potential and future avenues. Leukemia. 2011 Jun;25(6):909-20. https://doi.org/10.1038/leu.2011.48
- Marstrand TT, Borup R, Willer A, Borregaard N, Sandelin A, Porse BT, Theilgaard-Mönch K. A conceptual framework for the identification of candidate drugs and drug targets in acute promyelocytic leukemia. Leukemia. 2010 Jul;24(7):1265-75. https://www.nature.com/articles/le201095
- 2022 - Article by The Danish Cancer Society on new treatment with positive effect on patient: 'Jeg vil bare gerne have tid sammen med mine kære' (in Danish). See also this article by BRIC focused on the research: Translational AML trial improves clinical outcome and identifies novel drugs for future treatment
- 2020 – Lab member Montserrat Estruch reports on her recent AML publication in Leukemia: https://cancercommunity.nature.com/posts/targeted-inhibition-of-cooperative-mutation-and-therapy-induced-akt-activation-in-aml-effectively-enhances-response-to-chemotherapy
- 2019 - Research story about CD73 as a novel target in AML (Collaboration with Porse Group): https://www.bric.ku.dk/research-stories/research-stories-2019/study-points-to-cd73-as-novel-therapeutic-target-in-acute-myeloid-leukemia/
- 2018 - Research story on Kristian Reckzeh´s publication on the identification of a novel human blood stem cell marker: https://www.bric.ku.dk/research-stories/a-novel-blood-stem-cell-marker-for-improved-cancer-therapy/
- 2017 – Novo Nordisk Foundation highlighting the funding of the Program for Translational Hematology, a collaboration between scientists and clinicians at DanStem, Rigshospitalet, Finsen laboratory and BRIC: https://www.bric.ku.dk/research-stories/research-stories-2020/pth---towards-personalized-medicine/
- Olaf Heidenreich: Validation of novel ADCs in t(8;21) AML PDX models.
- Astra Zeneca: Validation of an Astra Zeneca second generation DNA-PK inhibitor in combination with standard chemotherapy in preclinical AML trials.
- Bo Porse, BRIC, UCPH, DK: CD73 as a novel therapeutic target for AML.
- Prof. Claus Storagard, BRIC, UCPH, DK: Validation of novel AML combination therapies including DNA-damage inhibitors and standard chemotherapy.
- Jesper Velgaard Olsen, CPR, UCPH, DK: AML proteomics.
- Lars Bullinger Charité, Berlin, DE: AML genetics.
- Clinical reseachers of Nordic and European AML trial groups (DK, NO, SE, FI, DE, NL)
- Novo Nordisk Foundation
- The Danish Cancer Society
- Independent Research Fund Denmark
- Lundbeck Foundation
- Børnecancerfonden
- Dagmar Marshalls Fond
- Dansk Kræftforskningsfond
- Læge Sofus Carl Emil Friis og Hustru Olga Doris Friis Foundation
- Tømrermester Jørgen Holm og Hustru Elisa
- Brødrene Hartmans Fond, F. Hansen's Mindelegat
- Sven Andersen Fonden
- iMED (EU PhD programme)
- Clinical Academic Group - UCPH
Conferences, Group dinners, Friday-Breakfasts, DHL race etc.
Group get together

Christmas card from Theilgaard-Mönch group


